👋 Hi I'm Flawnson

I'm a former ML engineer turned full-stack developer. I used to work a lot with Graph Neural Networks and NLP models. As a web developer I'm most competent in TS, React, Tailwind, Express, GraphQL, and Postgres.
I'm an avid piano player 🎹 and an abysmal frisbee tosser 🥏.
Given free time, I pander to and moderate the (small but growing) Geometric Deep Learning subreddit.
👨💻 What I'm working on
Most recently I've been working with my friend and co-founder Albert Wang on Comend. We build tools for the rare disease community to help them become research ready.
I recently started releasing original piano compositions on YouTube and Spotify.
For the latest updates on what I'm doing, my setup, or just to dig a bit more about me, visit my blog 📝.
📚 What I've worked on
Comend - Co-founder and CTO
Feb 2023 - Present - Toronto, Canada
- Built and launched Librarey, a search platform to help rare families find resources.
- Raised $600K in pre-seed funding to date from Character Labs and Drive Capital .
- Building Scimantic, a platform for generating plain language translations of academic and medical literature in the rare disease space with the goal of improving patient education and helping families build confidence to reach out to researchers.
Entrepreneur First (EF) - Founder in Residence
March 2022 - Feb 2023 - Toronto, Canada
- Had the honor of working alongside some of the brightest and most driven people I've met so far.
- Went through the formal process of finding strong co-founder fit and identifying problems worth solving and having useful customer conversations.
- Got very comfortable with sales tools like LinkedIn Sales Navigator, Apollo.io, Klenty, and Reply.io (kudos to those free trials).
Kebotix Inc. - Machine Learning Developer Intern
Sept 2020 - May 2021 - Boston, MA
- Developed a generative Transformer model optimized with a genetic algorithm for discovering organic and inorganic molecules of interest
- Contributing author of "Molecular discovery with conditional generative transformers coupled with genetic algorithms".
Relation Therapeutics - Machine Learning Research Intern
Sept 2019 - May 2020 - London, UK
- Developed novel GNN models for predicting the druggability and classifying the types of proteins.
- Built tuning and testing pipeline to optimize model parameters and run configurations.
- Contributing author of "Sparse Dynamic Distribution Decomposition: Efficient Integration of Trajectory and Snapshot Time Series Data".
ML for Chem/Bio-informatics - Personal Projects
Oct 2018 - June 2019 - Toronto, Canada
- Built a series of pipelines for preprocessing several chemical and biological data assays.
- Built various multivariate, evolutionary, predictive, and generative ML models to train on processed data.
- Wrote a series of articles on the various methods of Geometric Deep Learning.
Canadian Imperial Bank of Commerce (CIBC) - Front end Developer
June 2018 - Sept 2018 - Toronto, Canada
- Summer intern at CIBC Live Labs as front end developer and designer for an AR app.
- Designed basic UI/UX for app, collaborated with backend developers throughout development.
- Performed user/guerrilla testing of prototype and aggregated feedback for improvements.
📜 Publications
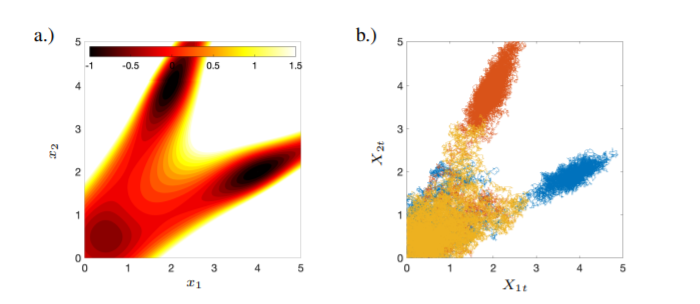
Sparse Dynamic Distribution Decomposition: Efficient Integration of Trajectory and Snapshot Time Series Data
Jake P. Taylor-King, Cristian Regep, Jyothish Soman, Flawnson Tong, Catalina Cangea, Charlie Roberts
DDD allows for the fitting of continuous-time Markov chains over these basis functions and as a result continuously maps between distributions. The number of parameters in DDD scales by the square of the number of basis functions; we reformulate the problem and restrict the method to compact basis functions which leads to the inference of sparse matrices only -- hence reducing the number of parameters.